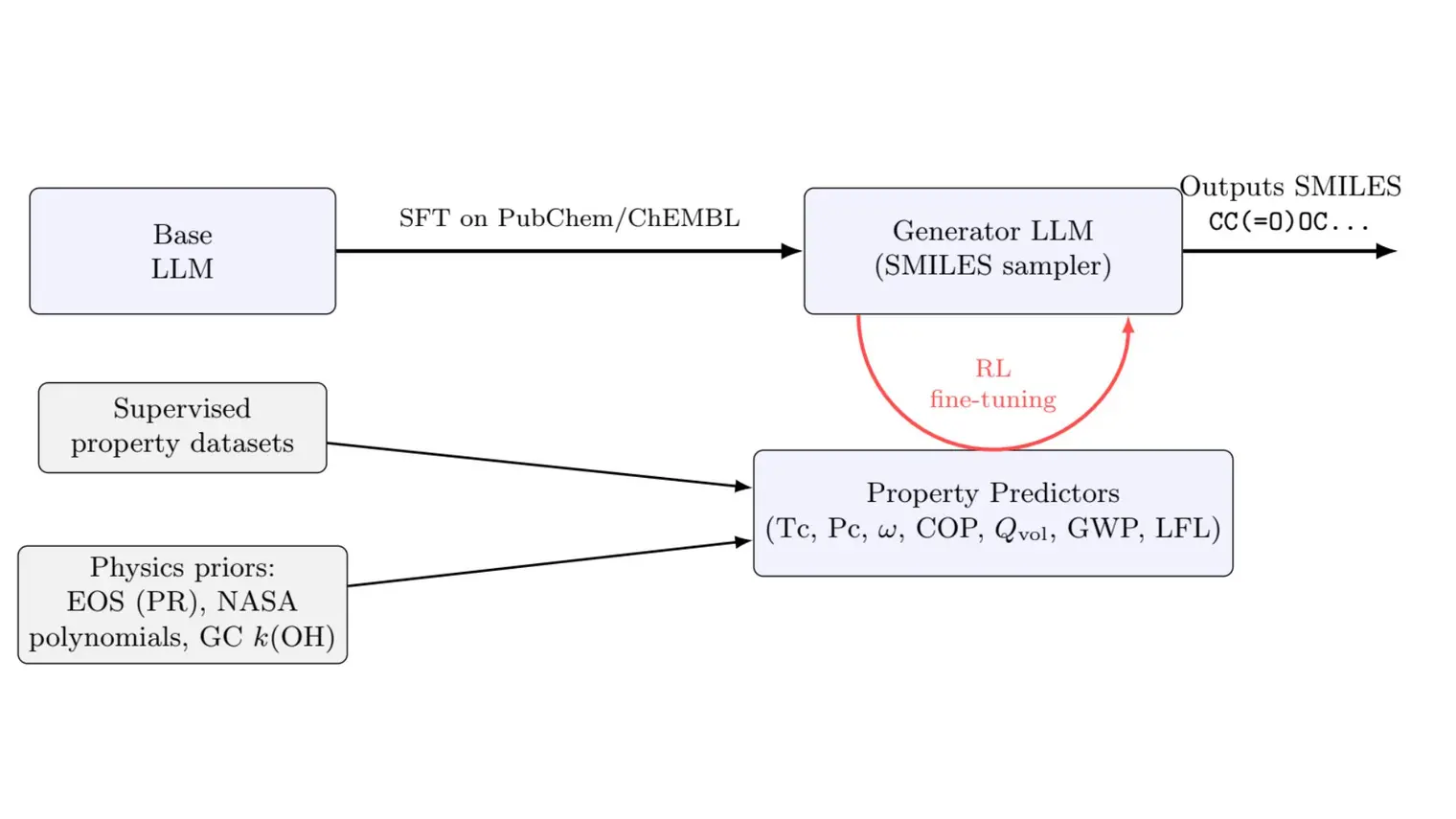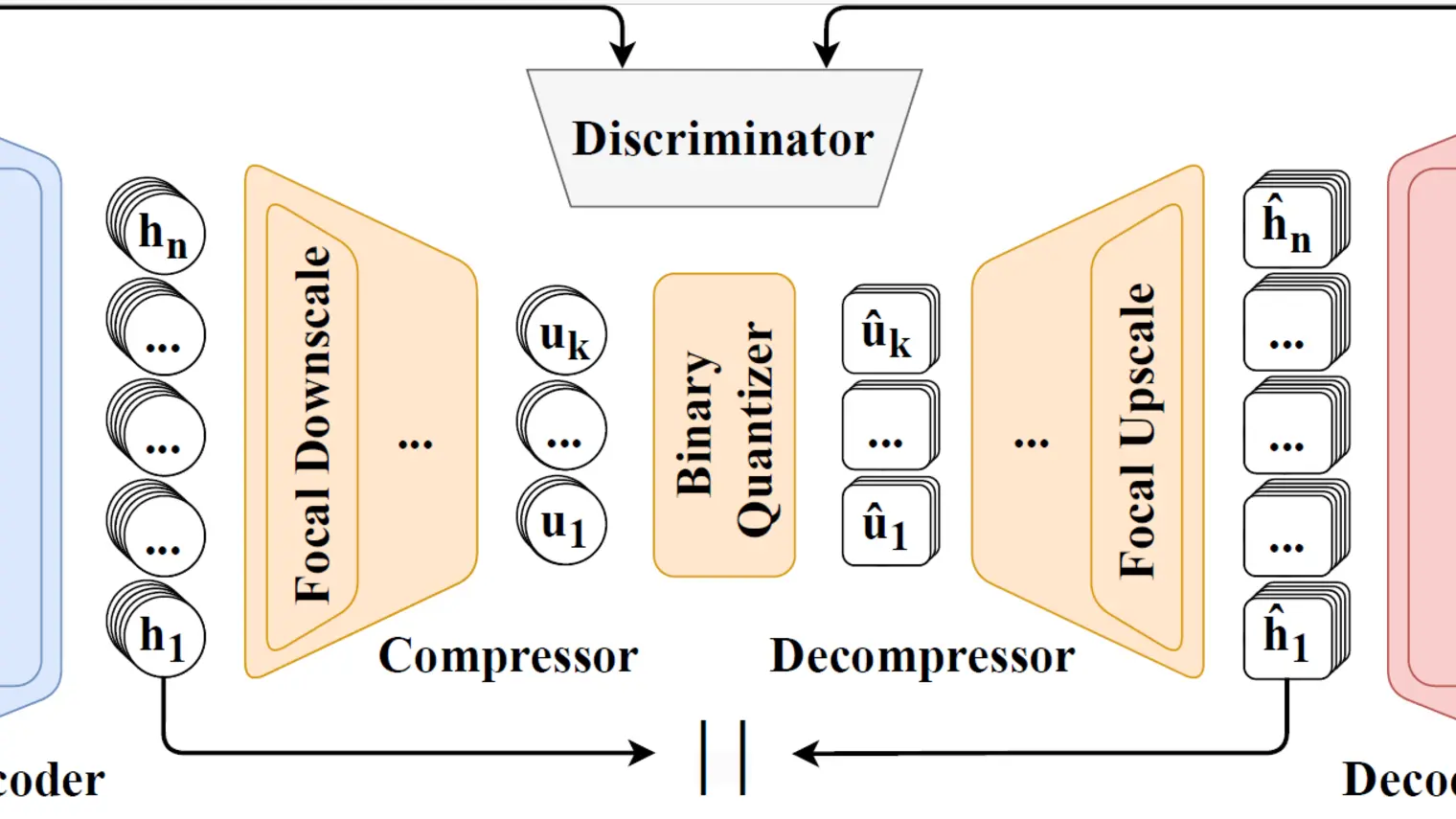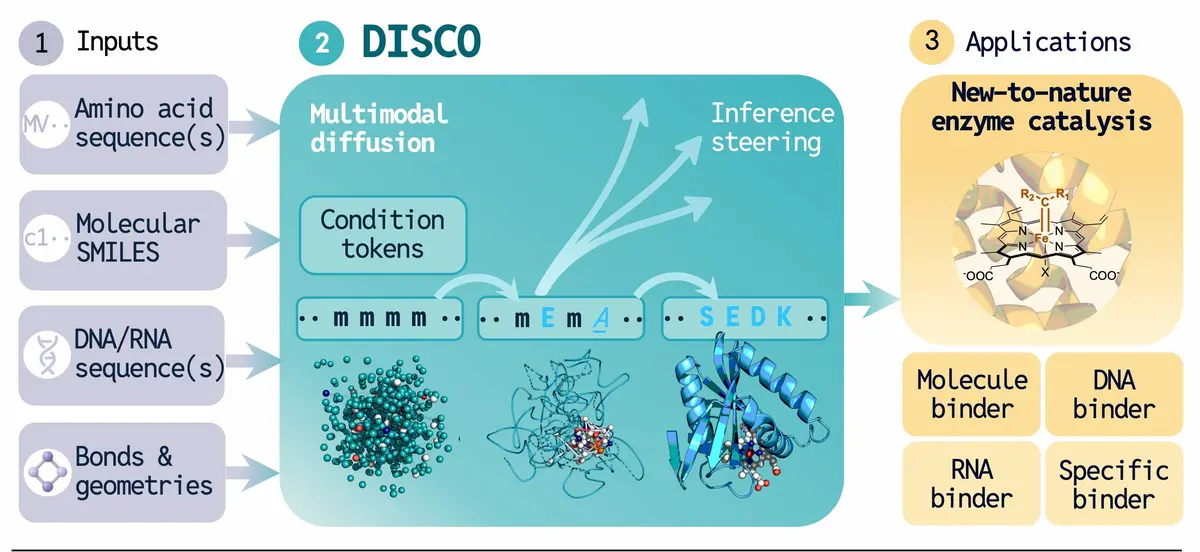
DISCO uses biological blueprints, like DNA or chemical structures, to simultaneously design a protein’s shape and its amino acid sequence. This allows scientists to create functional new-to-nature enzymes.
In 2016, a “plastic-eating” bacterium was discovered in a Japanese recycling facility. This ability comes from an enzyme, a natural molecular machine that can twist, break, and fuse molecules to create new substances.
Today, we can create new enzymes by mutating one that already exists in nature to give it a new function, called directed evolution. Yet this remains a time-consuming method limited by what nature has already invented.
Deep learning models have made spectacular advances in the de novo design of proteins. However, engineering new enzymes capable of catalyzing chemistry entirely unseen in nature remains a significant frontier.
Researchers from Mila, Caltech, Aithyra, FutureHouse and collaborators now introduce DISCO, a model that eliminates these bottlenecks.
A Single Model for Sequence, Structure, and Function
The shape of an enzyme is dictated by its amino acid sequence, which forces it to fold into a specific 3D structure. To function, specific amino acids must be strategically placed within this structure. Conventional models first generate the 3D shape and then attempt to find a matching sequence, but this disconnect fails to capture the complex interplay between chemistry and geometry. Furthermore, because current enzyme design methods rely on blueprints already found in nature, they struggle to invent new chemical functions.
Unlike traditional pipelines, DISCO simultaneously designs both the amino acid sequence and 3D structure, improving fidelity. This is enabled by:
- a unified multimodal loss: a single objective function that forces the model to optimize sequence and structure as an inseparable unit during training.
- a cross-modal recycling mechanism: an iterative feedback loop where sequence and structure data are continuously shared to ensure perfect mutual alignment.
- a self-correcting inference strategy: a real-time revision process that allows the model to detect and rectify inconsistencies during the enzyme generation process.

DISCO generates the most diverse, co-designable enzyme–ligand complexes for 178 of 179 targets in a new computational benchmark spanning diverse natural and non-natural molecules — outperforming all current baselines.
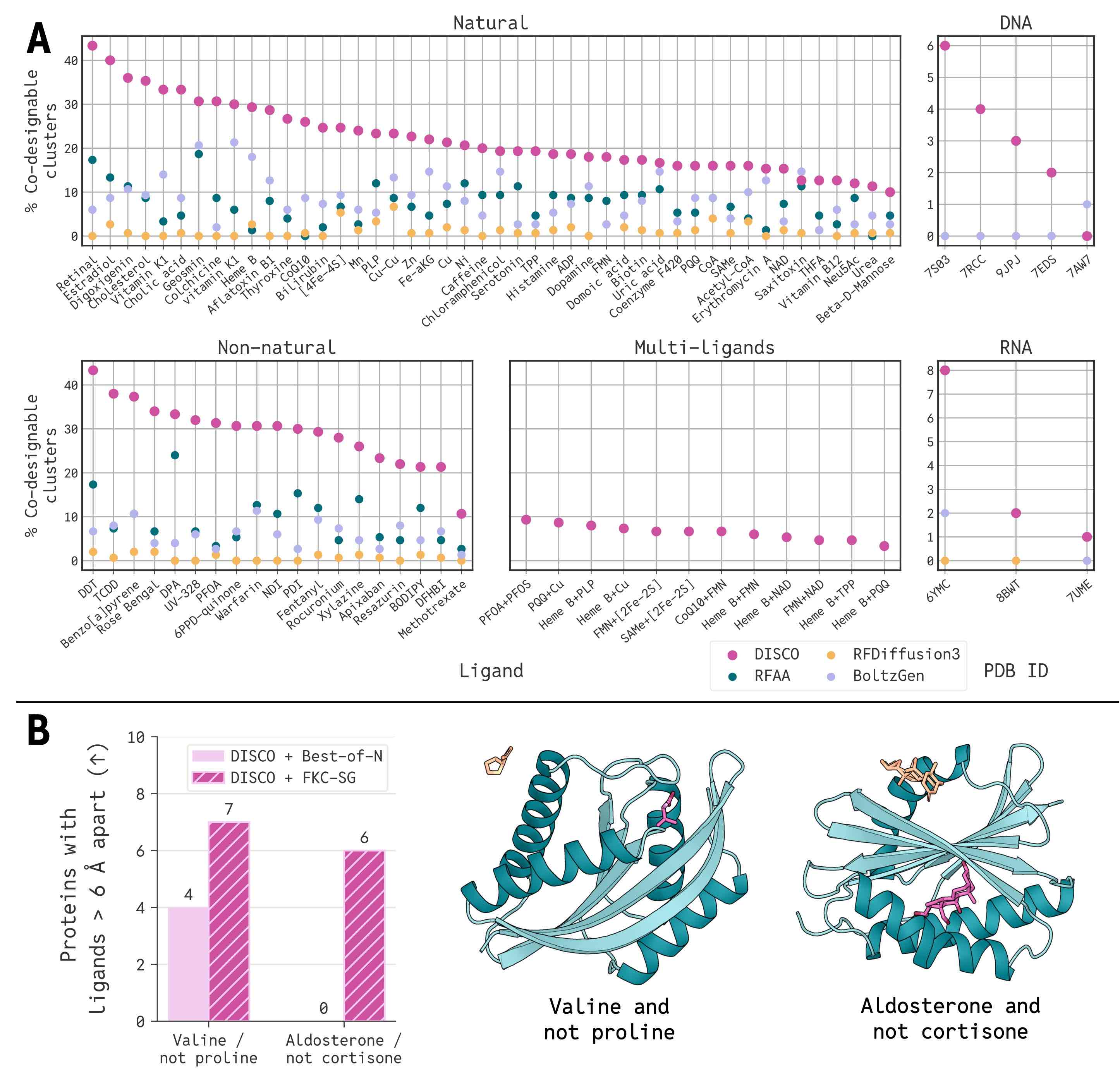
Designing Enzymes Without a Blueprint
90 enzyme designs generated by DISCO were synthesized in the laboratory and tested for their ability to catalyze 4 carbene-transfer reactions, a class of new-to-nature transformations that have proven valuable for constructing pharmaceutical drugs and complex molecules:
- B–H insertion: This reaction is entirely alien to biology. The best DISCO design achieved a 98% yield, with a single enzyme repeating the reaction 5,170 times, more than doubling the activity of previous enzymes selected through directed evolution.
- C(sp³)–H insertion: A difficult functionalization that reached 2,360 individual transformations per enzyme. This performance rivals results that previously required 14 successive rounds of laboratory-based evolution.
- Alkene cyclopropanation: This reaction achieved a 72% yield and over 4,000 reaction cycles per enzyme. It showed 99:1 diastereoselectivity (99% of the molecules produced have the exact same 3D atomic arrangement), surpassing the pioneer enzymes that first brought this chemistry into biology.
- Spirocyclopropanation: While initial activity was modest, a single round of random modifications quadrupled the enzyme's efficiency. It also enabled fine-grained control of the chirality of the final product. This rapid improvement suggests these designs are excellent starting points for further evolution.
Crucially, when compared against the AlphaFold Database of 200+ million structures, none of DISCO’s designed enzymes had active-site geometries with close natural homologs. DISCO didn't just remix known components; it invented functional molecular architectures that evolution never explored.
Project page: https://disco-design.github.io
Preprint: https://arxiv.org/abs/2604.05181